The quest to discover how diseases develop in humans has been hamstrung by high costs and access to large enough datasets. Scientists are racing to change that.
 Fully automated artificial intelligence could play a big role in advancing our understanding of how and why disease develop : NCI, Unsplash CC BY 4.0
Fully automated artificial intelligence could play a big role in advancing our understanding of how and why disease develop : NCI, Unsplash CC BY 4.0
The quest to discover how diseases develop in humans has been hamstrung by high costs and access to large enough datasets. Scientists are racing to change that.
It would be a eureka moment: discovering precisely how the development of diseases is linked to lifestyle, environment and genetics. But a huge amount of scientific research is needed to reach that milestone. Artificial intelligence (AI) is helping set the groundwork for the breakthrough, streamlining research with the goal of creating a fully automated system that can predict diseases and discover cures on its own.
In April 2022 Japan’s Okinawa Institute of Science and Technology (OIST) and the Tokyo-based Corundum Systems Biology Inc. began jointly working on a three-year project to establish an automated analytical system for microbiome and multi-omics data (data from various biological fields). The project is called MANTA: Multi-omics Analysis platform for the Nobel Turing challenge to develop AI scientists.
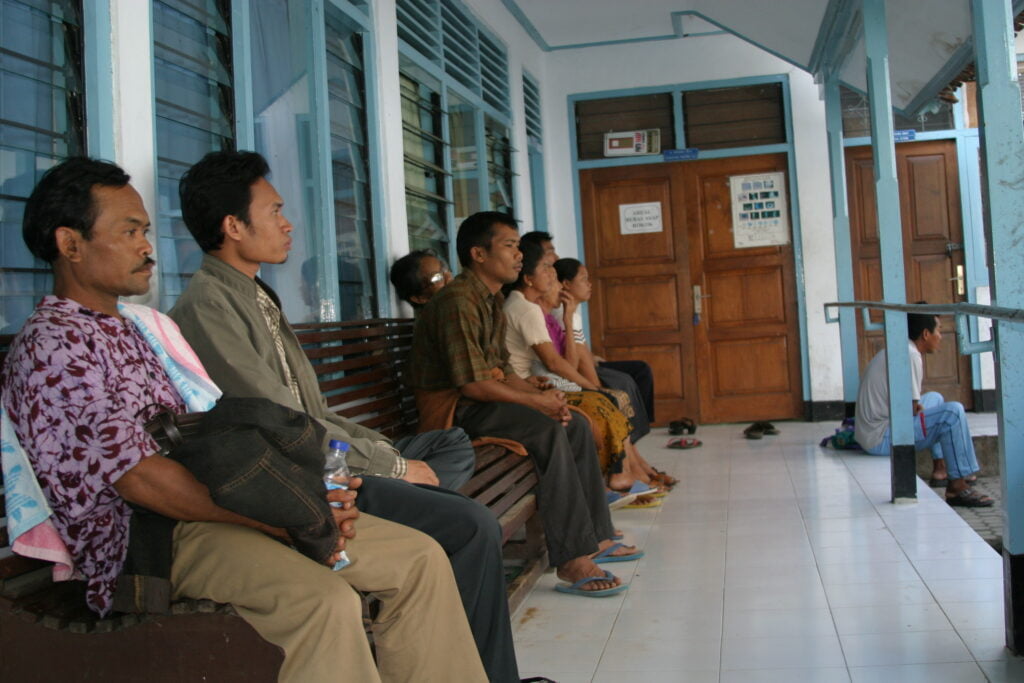
Alongside genetics, lifestyle factors such as dietary habits and sleep and environmental considerations such as exposure to pathogens and toxic substances are considered major determinants of human health. Getting to the bottom of this mix is complicated, so it requires epidemiological studies with large sample sizes to detect and quantify the impact of each factor.
To date, many projects, including case-control studies and prospective cohort studies for particular risk factors, have aimed to uncover the causes of disease.
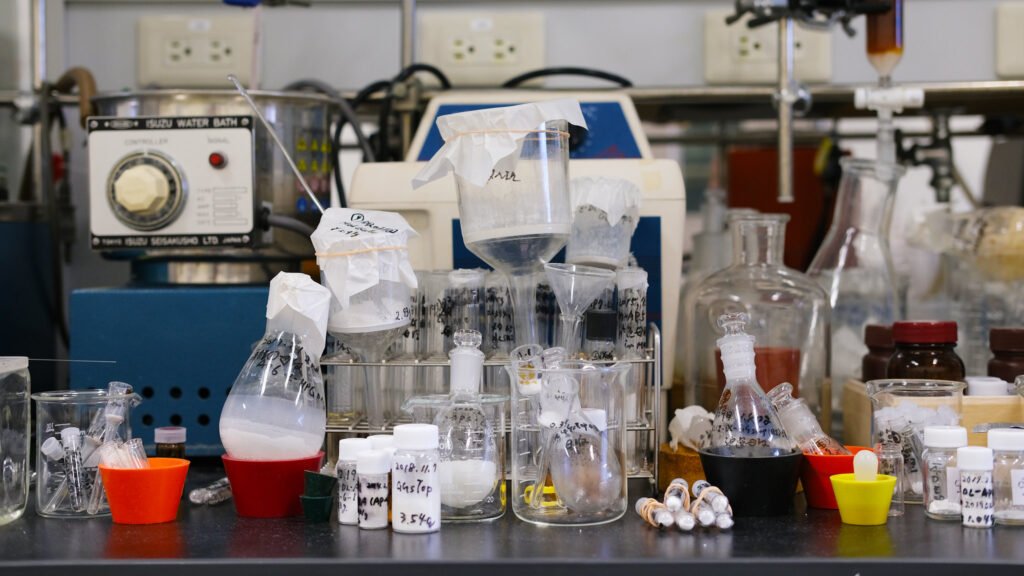
In many of these “phenotype” studies, biological samples such as blood, stool, urine and oral-swab samples are collected and tested. The resulting data guides further studies, linking a particular disease, lifestyle, location of residence or living environment with microbiome in many parts of the world. With more research, the involvement of the gut microbiome in the development and progression of multiple diseases, such as psychiatric disorders, metabolic diseases, brain diseases and cancer, gradually becomes clearer.
In the long term, the MANTA Project aims to better understand how living environments and factors unique to ethnic groups may be tied to causes of particular diseases. To date, plans to expand deep-phenotype study have been challenged by high costs and the time required to get the necessary biological data. Another major hurdle has been standardisation of data quality: the number of participants needed for each study is massive and the amount of data – from genomics, transcriptomics, proteomics and metabolomics to the microbiome of each study participant – is voluminous.
This makes AI-aided analysis critical to taking the next steps. Automated systems for gut microbiome and multi-omics data analysis can bring in globally consistent, replicable tests that can be reproduced en masse. It sets the stage for extensive data-building, as the same study participants can be monitored longer term and periodically at 10- to 20-year intervals, and at any location in the world. The automated nature of AI allows enormous groups of voluntary participants to be tracked for long periods, because it doesn’t involve the same cost and commitment of assigning humans to do the monitoring.
The goal and expectation is that in fully integrating AI into research about disease development, many unknowns will become known. What causes diseases to spread, worsen and change, and additional early signs or symptoms that research has yet to uncover, should reveal themselves with the new frontiers made possible by automated AI.
The MANTA Project is targeting full automation by March 2025, guiding a clear path for AI-supported analysis that – if successful – will speed up discoveries that assist both longevity and quality of life.
Dr Hiroaki Kitano is Professor (Adjunct) of Okinawa Institute of Science and Technology (OIST), Japan.
The MANTA project is partly funded by Corundum Systems Biology Inc, a Japanese company developing new businesses in the field of microbiome.
Originally published under Creative Commons by 360info™.